
Introducing daft.File: Work with Any File, Anywhere
daft.File brings lazy, distributed handling for audio, video, PDFs, and code to Daft DataFrames. One interface, local or remote.
by Everett KlevenDistributed Random Access for Audio, Video, Documents, and Code
Modern AI pipelines increasingly rely on unstructured files: audio, video, PDFs, source code, and more. These assets are often large, remote, and expensive to load eagerly.
daft.File brings file-native handling into Daft's lazy, distributed execution model. Instead of pulling full objects into memory, you can pass file references through your DataFrame, open them only when needed, and process them in parallel across your compute environment.
Since its introduction in Daft 0.6.0, daft.File has steadily matured into a full-featured capability. And with the recent additions of daft.VideoFile and daft.AudioFile, we're eager to see the community leverage daft.File for their own use-cases. If you run into any troubles please file an issue on github or reach out directly in our slack community.
Why daft.File
Daft already provides strong I/O performance for structured formats like Parquet, JSONL, and CSV. daft.File extends that same philosophy to unstructured data by giving you:
- Lazy file references in DataFrames
- Random-access reads through
open() - Disk materialization with
to_tempfile()for libraries that need a local path - Full reads with
read()when appropriate - Local and remote storage support with one interface
Basic Usage
Here's what that looks like in practice:
import daft
@daft.func
def read_header(f: daft.File) -> bytes:
with f.open() as fh:
return fh.read(16)
df = (
daft.from_glob_path("hf://datasets/Eventual-Inc/sample-files/")
.with_column("file", daft.functions.file(daft.col("path")))
.with_column("header", read_header(daft.col("file")))
.select("path", "size", "file", "header")
)
df.show(5)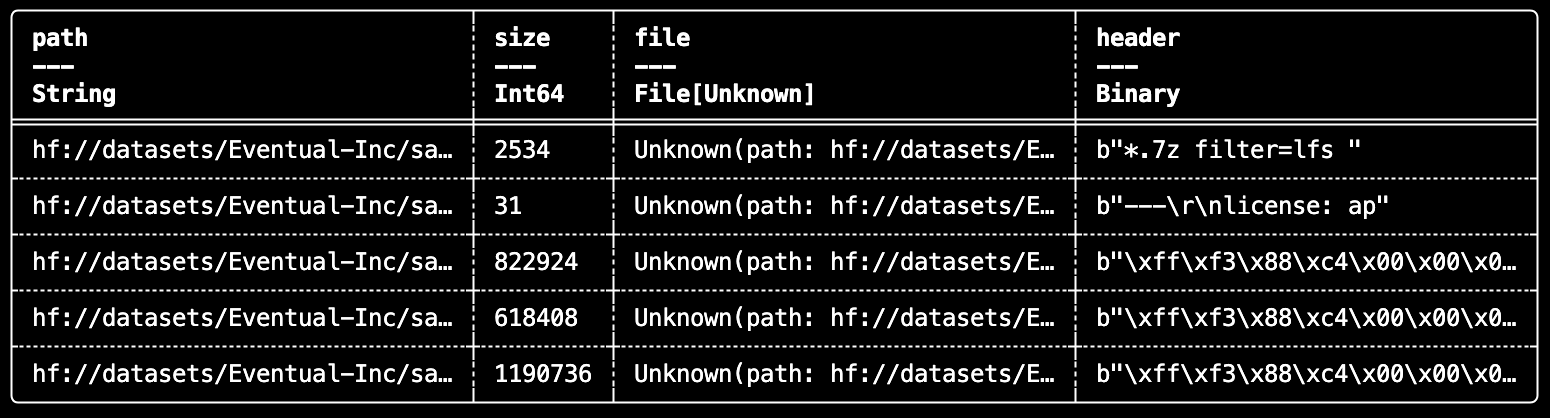
This pattern scales well because the file handle is only opened during execution, where it can run distributed across partitions.
Audio: Index, Read, and Resample with daft.AudioFile
With daft.AudioFile, you can index audio, inspect metadata, and prepare tensors for downstream ML tasks.
import daft
from daft.functions import audio_file, audio_metadata, resample
df = (
daft.from_glob_path("hf://datasets/Eventual-Inc/sample-files/audio/*.mp3")
.with_column("file", audio_file(daft.col("path")))
.with_column("metadata", audio_metadata(daft.col("file")))
.with_column("resampled", resample(daft.col("file"), sample_rate=16000))
.select("path", "file", "size", "metadata", "resampled")
)
df.show(3)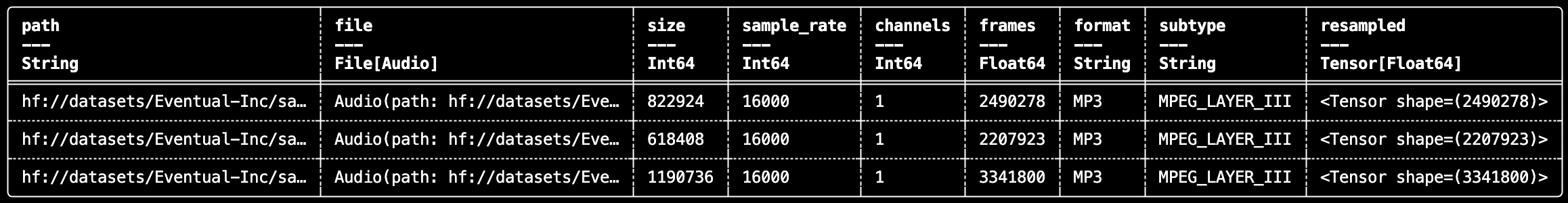
Video: Read Keyframes with daft.VideoFile
daft.VideoFile enables lightweight indexing and keyframe-aware video processing without writing custom ingestion glue.
import daft
from daft.functions import video_file, video_metadata, video_keyframes
df = (
daft.from_glob_path("hf://datasets/Eventual-Inc/sample-files/videos/*.mp4")
.with_column("file", video_file(daft.col("path")))
.with_column("metadata", video_metadata(daft.col("file")))
.with_column("keyframes", video_keyframes(daft.col("file")))
.select("path", "file", "size", "metadata", "keyframes")
)
df.show(3)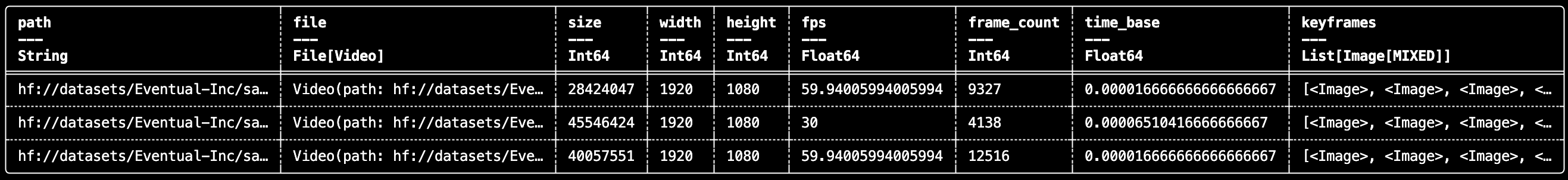
PDFs: Structured Extraction from Document Files
daft.File also works cleanly with document tooling. For example, you can convert each PDF to a tempfile and extract page-level text and rendered page images with PyMuPDF in a UDF.
import daft
import pymupdf
@daft.func(
return_dtype=daft.DataType.list(
daft.DataType.struct(
{
"page_number": daft.DataType.uint8(),
"page_text": daft.DataType.string(),
"page_image_bytes": daft.DataType.binary(),
}
)
)
)
def extract_pdf(file: daft.File):
"""Extracts the content of a PDF file."""
pymupdf.TOOLS.mupdf_display_errors(False) # Suppress non-fatal MuPDF warnings
content = []
with file.to_tempfile() as tmp:
doc = pymupdf.Document(filename=str(tmp.name), filetype="pdf")
for pno, page in enumerate(doc):
row = {
"page_number": pno,
"page_text": page.get_text("text"),
"page_image_bytes": page.get_pixmap().tobytes(),
}
content.append(row)
return content
if __name__ == "__main__":
from daft import col
df = (
daft.from_glob_path("hf://datasets/Eventual-Inc/sample-files/papers/*.pdf")
.with_column("pdf_file", daft.functions.file(col("path")))
.with_column("pages", extract_pdf(col("pdf_file")))
.explode("pages")
.select("path", "size", daft.functions.unnest(col("pages")))
)
df.show(3)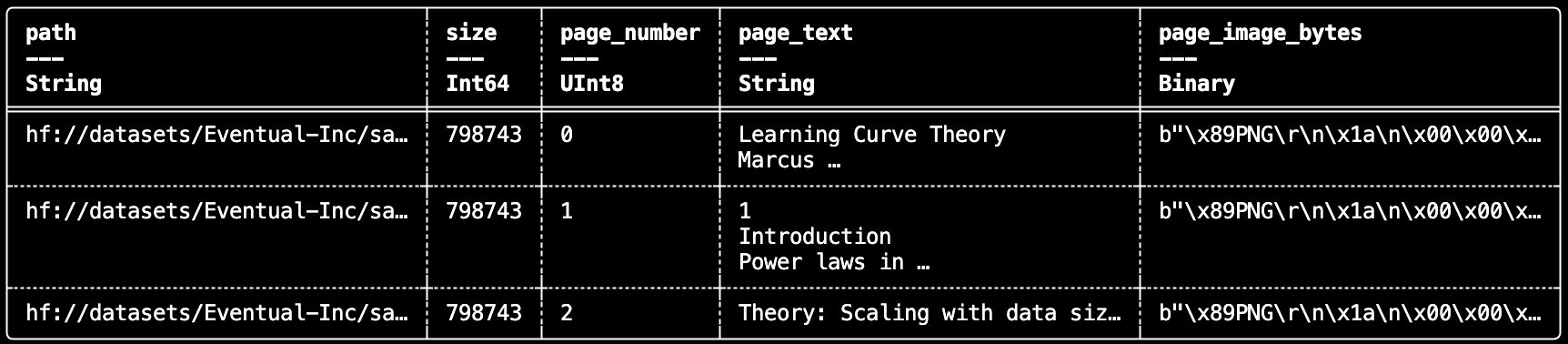
This gives you a straightforward way to build document ETL pipelines for retrieval, multimodal indexing, or downstream model input construction.
Code: Build Code Intelligence Pipelines with AST Parsing
Because daft.File is generic, the same interface works for source code analysis: open Python files, parse with ast, and extract function signatures/docstrings into structured rows. This is a strong pattern for code search, indexing, and repository analytics.
import daft
from daft import DataType, col
from daft.functions import unnest, file as daft_file
@daft.func(
return_dtype=DataType.list(
DataType.struct(
{
"name": DataType.string(),
"signature": DataType.string(),
"docstring": DataType.string(),
"start_line": DataType.int64(),
"end_line": DataType.int64(),
}
)
)
)
def extract_functions(file: daft.File):
"""Extract all function definitions from a Python file."""
import ast
with file.open() as f:
file_content = f.read().decode("utf-8")
tree = ast.parse(file_content)
results = []
for node in ast.walk(tree):
if isinstance(node, (ast.FunctionDef, ast.AsyncFunctionDef)):
signature = f"def {node.name}({ast.unparse(node.args)})"
if node.returns:
signature += f" -> {ast.unparse(node.returns)}"
results.append({
"name": node.name,
"signature": signature,
"docstring": ast.get_docstring(node),
"start_line": node.lineno,
"end_line": node.end_lineno,
})
return results
if __name__ == "__main__":
from daft import col
# Discover Python files from my local Daft Clone
df = (
daft.from_glob_path("~/git/Daft/daft/functions/**/*.py")
.with_column("file", daft_file(col("path")))
.with_column("functions", extract_functions(col("file")))
.explode("functions")
.select("path", "size", unnest(col("functions")))
)
df.show(3)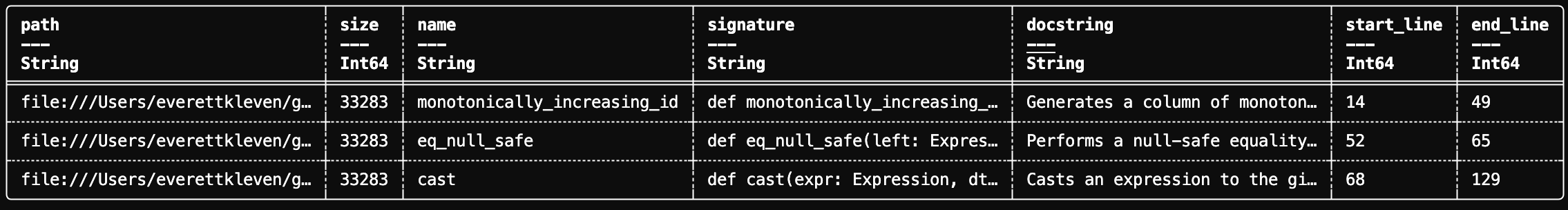
Unstructured Data, First-Class Treatment
daft.File bridges a major gap between tabular pipelines and real-world unstructured data. You can now treat files as first-class values in Daft: discover them, transform them lazily, and process them with Python/Rust ecosystem libraries at distributed scale.
If you're already using Daft for structured data, this gives you a direct path to unify audio, video, documents, and code in the same execution model.

